 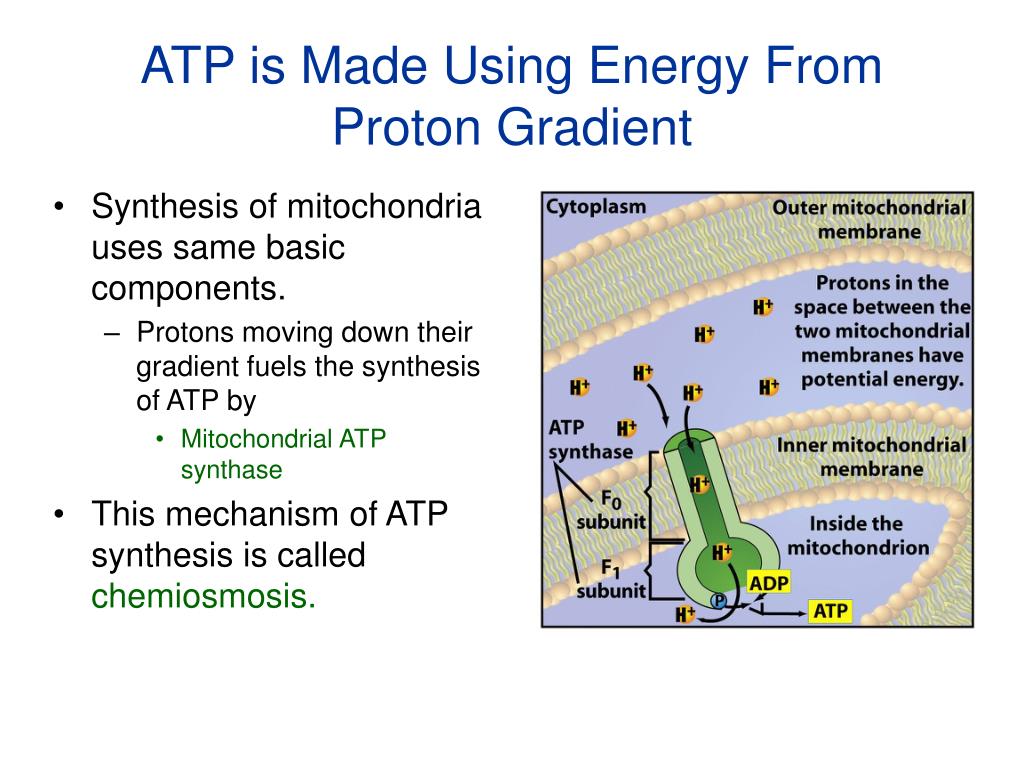
Crystal structure of bovine mitochondrial F1-ATPase with nucleotide bound to all three catalytic sites: implications for the mechanism of rotary catalysis. Observations of rotation within the FoF1-ATP synthase: deciding between rotation of the Fo c subunit ring and artifact. The striking direct demonstration of the rotation of the γ-subunit. Direct observation of the rotation of F1-ATPase. The demonstration of the molecular structure of the major part of the enzyme strongly indicated the rotation of the central γ-subunit surrounded by the cylinder of α 3 β 3 -subunits. 8 Å of F1-ATPase from bovine heart mitochondria. A gene cluster in the spinach and pea chloroplast genomes encoding one CF1 and three CFo subunits of the H +-ATP synthase complex and the ribosomal protein S2. Primary structure and subunit stoichiometry of F1-ATPase from bovine mitochondria. Nucleotide sequence of the genes coding for α-, β- and γ-subunits of the proton-translocating ATPase of Escherichia coli. This paper introduced a new concept of chemiosmotic theory into the field of bioenergetics. Coupling of phosphorylation to electron and hydrogen transfer by a chemiosmotic type mechanism. Boyer's rotational catalysis and alternate-binding change model are concisely reviewed. The ATP synthase - a splendid molecular machine. So, ATP synthase can be viewed as a complex of two motors: an ATP-driven F 1 motor and the proton-driven F o motor.īoyer, P. 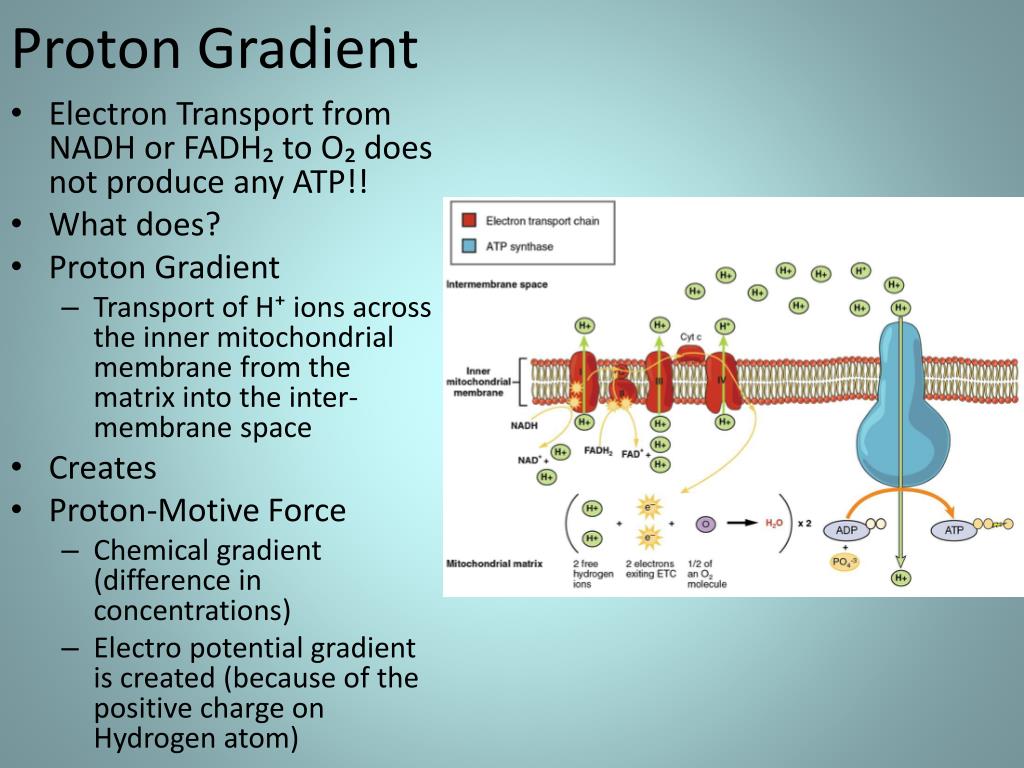
The synthesis of ATP is brought about by the rotary motion of the F oF 1 complex: when a large electrochemical potential (proton gradient) flows through the F o subunit, this causes rotation of the F o subunit and, subsequently, F 1, leading to ATP synthesis.ĪTP hydrolysis by ATPase - the reverse reaction - induces rotation of the F o rotor in the opposite direction. 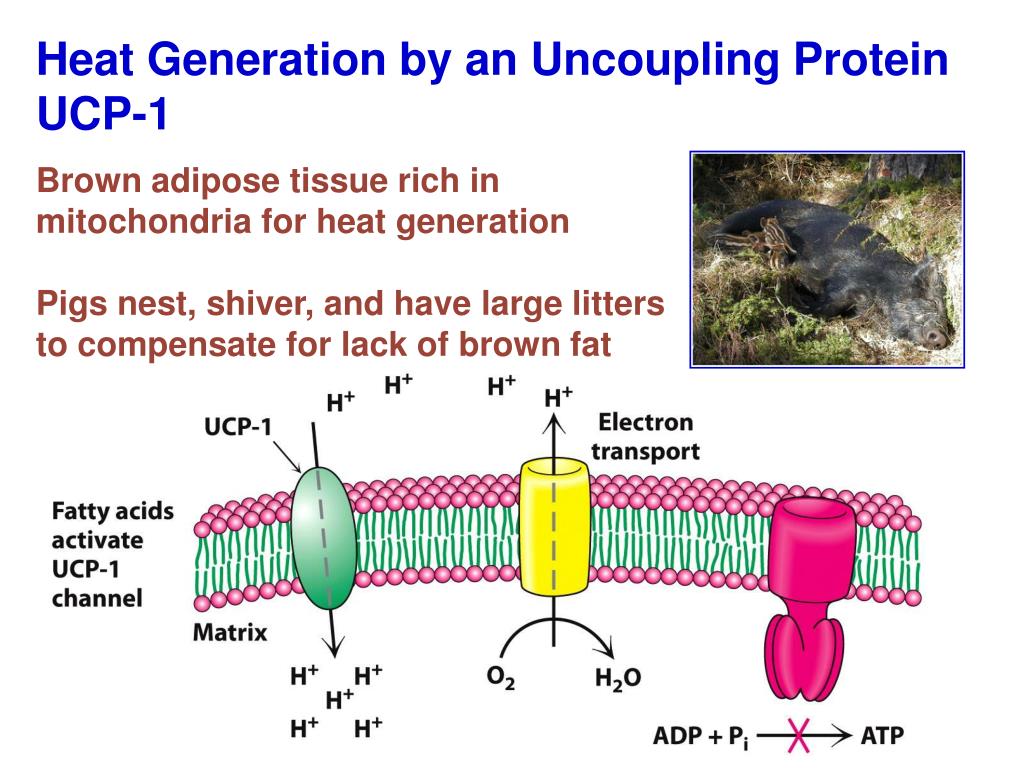
The F 1 portion consists of three α- and three β-subunits and a single γδɛ-subunit, whereas F o comprises one a-subunit, two b-subunits and 10–12 c-subunits. Recent analysis of the crystal structure of the enzyme has shown in atomic detail the intricate mechanisms of rotary catalysis.ĪTP synthase is a large (500 kDa) multisubunit protein, consisting of an intrinsic membrane domain, F o, linked through central and side stalks to a globular catalytic domain, F 1. The enzyme is located in the inner membrane of mitochondria, in the thylakoid membrane of chloroplasts, and in the plasma membrane of bacteria. As a result, the ATP synthase allows protons to re-enter the mitochondrial matrix and absorb their energy to produce ATP.ATP synthase is a ubiquitous, highly conserved enzyme that catalyses the formation of ATP from ADP and P i using a unique rotary motor mechanism. The F0 rotor spins when protons flow through the ATP synthase, facilitating the conversion of ADP and Pi to ATP. In ATP synthase, the "motor" is the F 0 subunit, whereas the "turbine" is the F 1 subunit, which uses a proton gradient to power ATP production. The proton gradient created by proton pumping during the electron transport chain is a confined energy source. The electrochemical proton gradient is a change in hydrogen ion concentration through a membrane that results in a concentration gradient with a slope and an electrical potential gradient. The proton gradient might be used to store energy for heat generation and flagella spinning. The oxidative phosphorylation process uses this potential energy to produce ATP. During photosynthesis and cellular respiration, the proton gradient is frequently employed to produce a chemiosmosis potential or proton motive force. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
